|
DMP_BBO library
|
|
DMP_BBO library
|
Base class for all function approximators. More...
#include <FunctionApproximator.hpp>


Public Member Functions | |
| FunctionApproximator (const MetaParameters *const meta_parameters, const ModelParameters *const model_parameters=NULL) | |
| Initialize a function approximator with meta- and optionally model-parameters. More... | |
| FunctionApproximator (const ModelParameters *const model_parameters) | |
| Initialize a function approximator with model-parameters. More... | |
| virtual FunctionApproximator * | clone (void) const =0 |
| Return a pointer to a deep copy of the FunctionApproximator object. More... | |
| virtual void | train (const Eigen::Ref< const Eigen::MatrixXd > &inputs, const Eigen::Ref< const Eigen::MatrixXd > &targets)=0 |
| Train the function approximator with corresponding input and target examples. More... | |
| void | train (const Eigen::Ref< const Eigen::MatrixXd > &inputs, const Eigen::Ref< const Eigen::MatrixXd > &targets, std::string save_directory, bool overwrite=false) |
| Train the function approximator with corresponding input and target examples (and write results to file). More... | |
| void | reTrain (const Eigen::Ref< const Eigen::MatrixXd > &inputs, const Eigen::Ref< const Eigen::MatrixXd > &targets) |
| Re-train the function approximator with corresponding input and target examples. More... | |
| void | reTrain (const Eigen::Ref< const Eigen::MatrixXd > &inputs, const Eigen::Ref< const Eigen::MatrixXd > &targets, std::string save_directory, bool overwrite=false) |
| Re-train the function approximator with corresponding input and target examples (and write results to file). More... | |
| virtual void | predict (const Eigen::Ref< const Eigen::MatrixXd > &inputs, Eigen::MatrixXd &outputs)=0 |
| Query the function approximator to make a prediction. More... | |
| virtual void | predict (const Eigen::Ref< const Eigen::MatrixXd > &inputs, Eigen::MatrixXd &outputs, Eigen::MatrixXd &variances) |
| Query the function approximator to make a prediction, and also to predict its variance. More... | |
| virtual void | predict (const Eigen::Ref< const Eigen::MatrixXd > &inputs, Eigen::MatrixXd &outputs, std::vector< Eigen::MatrixXd > &variances) |
| Query the function approximator to make a prediction, and also to predict its variance. More... | |
| virtual void | predictVariance (const Eigen::Ref< const Eigen::MatrixXd > &inputs, Eigen::MatrixXd &variances) |
| Query the function approximator to get the variance of a prediction This function is not implemented by all function approximators. More... | |
| bool | isTrained (void) const |
| Determine whether the function approximator has already been trained with data or not. More... | |
| int | getExpectedInputDim (void) const |
| The expected dimensionality of the input data. More... | |
| int | getExpectedOutputDim (void) const |
| The expected dimensionality of the output data. More... | |
| virtual std::string | getName (void) const =0 |
| Get the name of this function approximator. More... | |
| void | getSelectableParameters (std::set< std::string > &selected_values_labels) const |
| Return all the names of the parameter types that can be selected. More... | |
| void | setSelectedParameters (const std::set< std::string > &selected_values_labels) |
| Determine which subset of parameters is represented in the vector returned by Parameterizable::getParameterVectorSelected. More... | |
| void | getParameterVectorSelectedMinMax (Eigen::VectorXd &min, Eigen::VectorXd &max) const |
| Get the minimum and maximum of the selected parameters in one vector. More... | |
| int | getParameterVectorSelectedSize (void) const |
| Get the size of the vector of selected parameters, as returned by getParameterVectorSelected(. More... | |
| void | setParameterVectorSelected (const Eigen::VectorXd &values, bool normalized=false) |
| Set all the values of the selected parameters with one vector. More... | |
| void | getParameterVectorSelected (Eigen::VectorXd &values, bool normalized=false) const |
| Get the values of the selected parameters in one vector. More... | |
| void | getParameterVectorMask (const std::set< std::string > selected_values_labels, Eigen::VectorXi &selected_mask) const |
| Get a mask for selecting parameters. More... | |
| int | getParameterVectorAllSize (void) const |
| Get the size of the parameter values vector when it contains all available parameter values. More... | |
| void | getParameterVectorAll (Eigen::VectorXd &values) const |
| Return a vector that returns all available parameter values. More... | |
| void | setParameterVectorAll (const Eigen::VectorXd &values) |
| Set all available parameter values with one vector. More... | |
| UnifiedModel * | getUnifiedModel (void) const |
| Return a representation of this function approximator's model as a unified model. More... | |
| std::string | toString (void) const |
| Returns a string representation of the object. More... | |
| const MetaParameters * | getMetaParameters (void) const |
| Accessor for FunctionApproximator::meta_parameters_. More... | |
| const ModelParameters * | getModelParameters (void) const |
| Accessor for FunctionApproximator::model_parameters_. More... | |
| void | setParameterVectorModifierPrivate (std::string modifier, bool new_value) |
| Turn certain modifiers on or off, see Parameterizable::setParameterVectorModifier(). More... | |
| virtual bool | saveGridData (const Eigen::VectorXd &min, const Eigen::VectorXd &max, const Eigen::VectorXi &n_samples_per_dim, std::string directory, bool overwrite=false) const |
| Generate a grid of inputs, and output the response of the basis functions and line segments for these inputs. More... | |
 Public Member Functions inherited from Parameterizable Public Member Functions inherited from Parameterizable | |
| virtual | ~Parameterizable (void) |
| Destructor. | |
| virtual void | getParameterVectorSelectedNormalized (Eigen::VectorXd &values) const |
| Get the normalized values of the selected parameters in one vector. More... | |
| void | getParameterVectorSelectedMinMax (Eigen::VectorXd &min, Eigen::VectorXd &max) const |
| Get the minimum and maximum of the selected parameters in one vector. More... | |
| void | getParameterVectorAllMinMax (Eigen::VectorXd &min, Eigen::VectorXd &max) const |
| Get the minimum and maximum values of the current parameter vector. More... | |
| void | getParameterVectorSelectedRanges (Eigen::VectorXd &ranges) const |
| Get the ranges of the selected parameters, i.e. More... | |
| virtual void | setParameterVectorSelectedNormalized (const Eigen::VectorXd &values) |
| Set all the values of the selected parameters with one vector of normalized values. More... | |
| void | setSelectedParametersOne (std::string selected) |
| Set the parameters that are currently selected. More... | |
| void | setParameterVectorModifier (std::string modifier, bool new_value) |
| Turn certain modifiers on or off. More... | |
| void | setVectorLengthsPerDimension (const Eigen::VectorXi &lengths_per_dimension) |
| The vector (VectorXd) with parameter values can be split into different parts (as vector<VectorXd>; this function specifices the length of each sub-vector. More... | |
| Eigen::VectorXi | getVectorLengthsPerDimension (void) const |
| Get the specified length of each vector in each dimension. More... | |
| void | getParameterVectorSelected (std::vector< Eigen::VectorXd > &values, bool normalized=false) const |
| Get the values of the selected parameters in one vector. More... | |
| void | setParameterVectorSelected (const std::vector< Eigen::VectorXd > &values, bool normalized=false) |
| Set all the values of the selected parameters with a vector of vectors. More... | |
Static Public Member Functions | |
| static void | generateInputsGrid (const Eigen::VectorXd &min, const Eigen::VectorXd &max, const Eigen::VectorXi &n_samples_per_dim, Eigen::MatrixXd &inputs_grid) |
| Generate a input samples that lie on a grid (much like Matlab's meshgrid) For instance, if min = [2 6], and max = [3 8], and n_samples_per_dim = [3 5] then this function first makes linearly spaces samples along each dimension, e.g. More... | |
Protected Member Functions | |
| void | setModelParameters (ModelParameters *model_parameters) |
| Accessor for FunctionApproximator::model_parameters_. More... | |
| FunctionApproximator (void) | |
| Default constructor. More... | |
Friends | |
| class | boost::serialization::access |
| Give boost serialization access to private members. More... | |
| std::ostream & | operator<< (std::ostream &output, const FunctionApproximator &function_approximator) |
| Print to output stream. More... | |
Base class for all function approximators.
Definition at line 48 of file FunctionApproximator.hpp.
| FunctionApproximator | ( | const MetaParameters *const | meta_parameters, |
| const ModelParameters *const | model_parameters = NULL |
||
| ) |
Initialize a function approximator with meta- and optionally model-parameters.
| [in] | meta_parameters | The training algorithm meta-parameters |
| [in] | model_parameters | The parameters of the trained model. If this parameter is not passed, the function approximator is initialized as untrained. In this case, you must call FunctionApproximator::train() before being able to call FunctionApproximator::predict(). Either meta_parameters XOR model-parameters can passed as NULL, but not both. |
Definition at line 49 of file FunctionApproximator.cpp.

| FunctionApproximator | ( | const ModelParameters *const | model_parameters | ) |
Initialize a function approximator with model-parameters.
| [in] | model_parameters | The parameters of the trained model. |
Definition at line 70 of file FunctionApproximator.cpp.

|
inlineprotected |
Default constructor.
Definition at line 308 of file FunctionApproximator.hpp.
|
pure virtual |
Return a pointer to a deep copy of the FunctionApproximator object.
Implemented in FunctionApproximatorRRRFF, FunctionApproximatorLWPR, FunctionApproximatorGMR, FunctionApproximatorGPR, FunctionApproximatorLWR, and FunctionApproximatorRBFN.
|
pure virtual |
Train the function approximator with corresponding input and target examples.
| [in] | inputs | Input values of the training examples |
| [in] | targets | Target values of the training examples |
Implemented in FunctionApproximatorRRRFF, FunctionApproximatorLWPR, FunctionApproximatorGMR, FunctionApproximatorGPR, FunctionApproximatorLWR, and FunctionApproximatorRBFN.
| void train | ( | const Eigen::Ref< const Eigen::MatrixXd > & | inputs, |
| const Eigen::Ref< const Eigen::MatrixXd > & | targets, | ||
| std::string | save_directory, | ||
| bool | overwrite = false |
||
| ) |
Train the function approximator with corresponding input and target examples (and write results to file).
| [in] | inputs | Input values of the training examples |
| [in] | targets | Target values of the training examples |
| [in] | save_directory | Directory to which to write results. |
| [in] | overwrite | Whether to overwrite existing files. true=do overwrite, false=don't overwrite and give a warning. |
Definition at line 288 of file FunctionApproximator.cpp.
| void reTrain | ( | const Eigen::Ref< const Eigen::MatrixXd > & | inputs, |
| const Eigen::Ref< const Eigen::MatrixXd > & | targets | ||
| ) |
Re-train the function approximator with corresponding input and target examples.
| [in] | inputs | Input values of the training examples |
| [in] | targets | Target values of the training examples Re-training could in principle have been enabled through FunctionApproximator::train, but we wanted to keep a clear disctinction between training (which must be done at least once before FunctionApproximator::predict) can be called and re-training. |
Definition at line 130 of file FunctionApproximator.cpp.
| void reTrain | ( | const Eigen::Ref< const Eigen::MatrixXd > & | inputs, |
| const Eigen::Ref< const Eigen::MatrixXd > & | targets, | ||
| std::string | save_directory, | ||
| bool | overwrite = false |
||
| ) |
Re-train the function approximator with corresponding input and target examples (and write results to file).
| [in] | inputs | Input values of the training examples |
| [in] | targets | Target values of the training examples |
| [in] | save_directory | Directory to which to write results. |
| [in] | overwrite | Overwrite existing files in the directory above (default: false) Re-training could in principle have been enabled through FunctionApproximator::train, but we wanted to keep a clear disctinction between training (which must be done at least once before FunctionApproximator::predict) can be called and re-training. |
Definition at line 137 of file FunctionApproximator.cpp.
|
pure virtual |
Query the function approximator to make a prediction.
| [in] | inputs | Input values of the query |
| [out] | outputs | Predicted output values |
Implemented in FunctionApproximatorLWR, FunctionApproximatorRRRFF, FunctionApproximatorGMR, FunctionApproximatorGPR, FunctionApproximatorLWPR, and FunctionApproximatorRBFN.
|
inlinevirtual |
Query the function approximator to make a prediction, and also to predict its variance.
| [in] | inputs | Input values of the query (n_samples X n_dims_in) |
| [out] | outputs | Predicted output values (n_samples X n_dims_out) |
| [out] | variances | Predicted variances for the output values (n_samples X n_dims_out). Note that if the output has a dimensionality>1, these variances should actuall be covariance matrices (use function predict(const Eigen::Ref<const Eigen::MatrixXd>& inputs, Eigen::MatrixXd& outputs, std::vector<Eigen::MatrixXd>& variances) to get the full covariance matrices). So for an output dimensionality of 1 this function works fine. For dimensionality>1 we return only the diagional of the covariance matrix, which may not always be what you want. |
Reimplemented in FunctionApproximatorGMR.
Definition at line 142 of file FunctionApproximator.hpp.

|
inlinevirtual |
Query the function approximator to make a prediction, and also to predict its variance.
| [in] | inputs | Input values of the query (n_samples X n_dims_in) |
| [out] | outputs | Predicted output values (n_samples X n_dims_out) |
| [out] | variances | Predicted covariance matrices for the output values. It is is of size (n_samples X n_dims_out X n_dims_out), which has been implemented as a std::vector of Eigen::MatrixXd. |
Definition at line 160 of file FunctionApproximator.hpp.

|
inlinevirtual |
Query the function approximator to get the variance of a prediction This function is not implemented by all function approximators.
Therefore, the default implementation fills outputs with 0s.
| [in] | inputs | Input values of the query (n_samples X n_dims_in) |
| [out] | variances | Predicted variances for the output values (n_samples X n_dims_out). Note that if the output has a dimensionality>1, these variances should actuall be covariance matrices (use function predict(const Eigen::Ref<const Eigen::MatrixXd>& inputs, Eigen::MatrixXd& outputs, std::vector<Eigen::MatrixXd>& variances) to get the full covariance matrices). So for an output dimensionality of 1 this function works fine. For dimensionality>1 we return only the diagional of the covariance matrix, which may not always be what you want. |
Reimplemented in FunctionApproximatorGMR, and FunctionApproximatorGPR.
Definition at line 181 of file FunctionApproximator.hpp.
|
inline |
Determine whether the function approximator has already been trained with data or not.
Definition at line 191 of file FunctionApproximator.hpp.
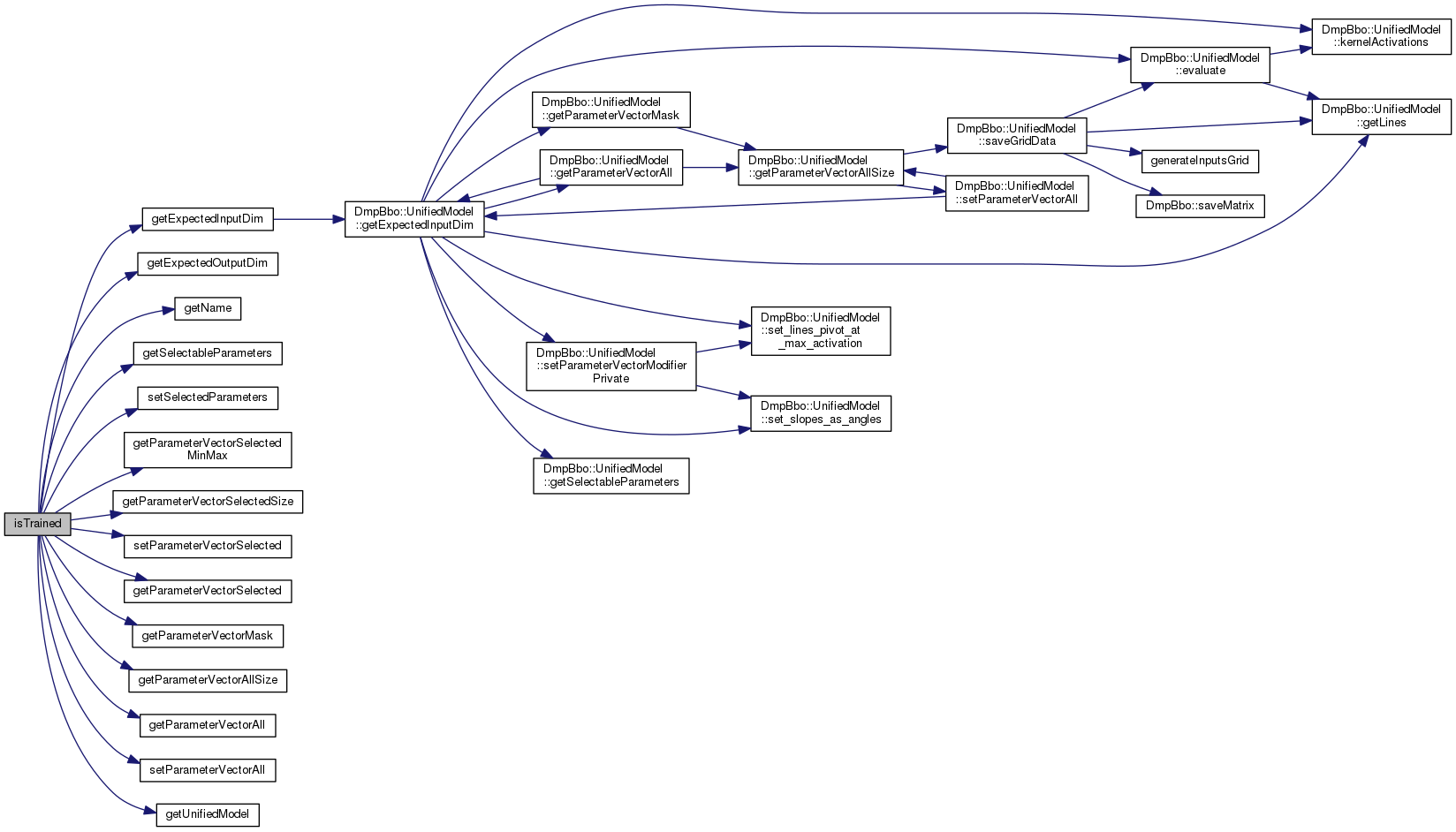
| int getExpectedInputDim | ( | void | ) | const |
The expected dimensionality of the input data.
Definition at line 114 of file FunctionApproximator.cpp.

| int getExpectedOutputDim | ( | void | ) | const |
The expected dimensionality of the output data.
Definition at line 122 of file FunctionApproximator.cpp.
|
pure virtual |
Get the name of this function approximator.
Implemented in FunctionApproximatorLWR, FunctionApproximatorRBFN, FunctionApproximatorGMR, FunctionApproximatorGPR, FunctionApproximatorRRRFF, and FunctionApproximatorLWPR.
|
virtual |
Return all the names of the parameter types that can be selected.
| [out] | selected_values_labels | Names of the parameter types that can be selected |
Implements Parameterizable.
Definition at line 200 of file FunctionApproximator.cpp.
|
virtual |
Determine which subset of parameters is represented in the vector returned by Parameterizable::getParameterVectorSelected.
Different function approximators have different types of model parameters. For instance, LWR has the centers and widths of basis functions, along with the slopes of each line segment. Parameterizable::setSelectedParameters provides a means to determine which parameters should be returned by Parameterizable::getParameterVectorSelected, i.e. by calling: std::set<std::string> selected; selected.insert("slopes"); model_parameters.setSelectedParameters(selected)
| [in] | selected_values_labels | The names of the parameters that are selected |
Reimplemented from Parameterizable.
Definition at line 194 of file FunctionApproximator.cpp.
| void getParameterVectorSelectedMinMax | ( | Eigen::VectorXd & | min, |
| Eigen::VectorXd & | max | ||
| ) | const |
Get the minimum and maximum of the selected parameters in one vector.
| [out] | min | The minimum of the selected parameters concatenated in one vector |
| [out] | max | The minimum of the selected parameters concatenated in one vector |
Definition at line 145 of file FunctionApproximator.cpp.
|
virtual |
Get the size of the vector of selected parameters, as returned by getParameterVectorSelected(.
Reimplemented from Parameterizable.
Definition at line 180 of file FunctionApproximator.cpp.
|
virtual |
Set all the values of the selected parameters with one vector.
| [in] | values | The new values of the selected parameters in one vector |
| [in] | normalized | Whether the data is normalized or not |
Reimplemented from Parameterizable.
Definition at line 188 of file FunctionApproximator.cpp.
|
virtual |
Get the values of the selected parameters in one vector.
| [out] | values | The selected parameters concatenated in one vector |
| [in] | normalized | Whether to normalize the data or not |
Reimplemented from Parameterizable.
Definition at line 171 of file FunctionApproximator.cpp.
|
virtual |
Get a mask for selecting parameters.
| [in] | selected_values_labels | Labels of the selected parameter values |
| [out] | selected_mask | A mask indicating indices of selected parameters. 0 indicates not selected, >0 indicates selected. |
For instance, if the parameters consists of centers, widths and slopes the parameter values vector will be something like
centers widths slopes [ 100 110 120 10 10 10 0.4 0.7 0.4 ]
In this case, if selected_values_labels contains "centers" and "slopes", the mask will be:
centers widths slopes [ 1 1 1 0 0 0 3 3 3 ]
The '0' indicates that these parameters are not selected. The other ones have different numbers so that they may be discerned from one another (as required in Parameterizable::getParameterVectorSelectedMinMax for instance.
Implements Parameterizable.
Definition at line 208 of file FunctionApproximator.cpp.
|
virtual |
Get the size of the parameter values vector when it contains all available parameter values.
For instance, if the parameters consists of centers, widths and slopes the parameter values vector will be something like
centers widths slopes [ 100 110 120 10 10 10 0.4 0.7 0.4 ]
then getParameterVectorAllSize() will return 9
Implements Parameterizable.
Definition at line 215 of file FunctionApproximator.cpp.
|
virtual |
Return a vector that returns all available parameter values.
| [out] | values | All available parameter values in one vector. |
Implements Parameterizable.
Definition at line 221 of file FunctionApproximator.cpp.
|
virtual |
Set all available parameter values with one vector.
| [in] | values | All available parameter values in one vector. |
Implements Parameterizable.
Definition at line 227 of file FunctionApproximator.cpp.
| UnifiedModel * getUnifiedModel | ( | void | ) | const |
Return a representation of this function approximator's model as a unified model.
See also the page on Unified Model for Function Approximators
Definition at line 108 of file FunctionApproximator.cpp.
| string toString | ( | void | ) | const |
Returns a string representation of the object.
Definition at line 232 of file FunctionApproximator.cpp.
| const MetaParameters * getMetaParameters | ( | void | ) | const |
Accessor for FunctionApproximator::meta_parameters_.
Definition at line 85 of file FunctionApproximator.cpp.
| const ModelParameters * getModelParameters | ( | void | ) | const |
Accessor for FunctionApproximator::model_parameters_.
Definition at line 91 of file FunctionApproximator.cpp.
|
virtual |
Turn certain modifiers on or off, see Parameterizable::setParameterVectorModifier().
Parameterizable::setParameterVectorModifierPrivate(), This function may (but must not be) overridden by subclasses of Parameterizable, depending on whether the subclass has modifiers (or not)
| [in] | modifier | The name of the modifier |
| [in] | new_value | Whether to turn the modifier on (true) or off (false) |
Reimplemented from Parameterizable.
Definition at line 390 of file FunctionApproximator.cpp.
|
static |
Generate a input samples that lie on a grid (much like Matlab's meshgrid) For instance, if min = [2 6], and max = [3 8], and n_samples_per_dim = [3 5] then this function first makes linearly spaces samples along each dimension, e.g.
samples_dim1 = [2 2.5 3] and samples_dim2 = [6 6.5 7 7.5 8] and then combine the combination of samples along all dimensions: inputs_grid = [2 6; 2 6.5; 2 7; 2 7.5 2 8; 2.5 6; 2.5 6.5; 2.5 7; 2.5 7.5 2.5 8; 3 6; 3 6.5; 3 7; 3 7.5 3 8;
| [in] | min | Minimum values in the grid along each dimension |
| [in] | max | Maximum values in the grid along each dimension |
| [in] | n_samples_per_dim | Number of samples along each dimension in the grid |
| [out] | inputs_grid | A grid of samples |
Definition at line 238 of file FunctionApproximator.cpp.
|
virtual |
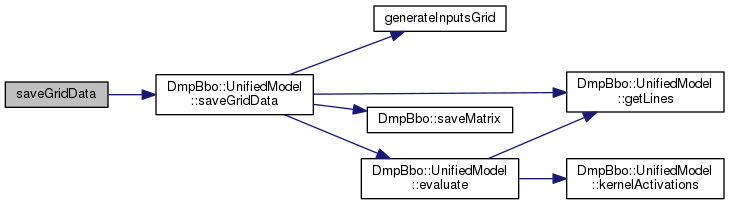
Generate a grid of inputs, and output the response of the basis functions and line segments for these inputs.
This function is not pure virtual, because this might not make sense for every model parameters class.
| [in] | min | Minimum values for the grid (one for each dimension) |
| [in] | max | Maximum values for the grid (one for each dimension) |
| [in] | n_samples_per_dim | Number of samples in the grid along each dimension |
| [in] | directory | Directory to which to save the results to. |
| [in] | overwrite | Whether to overwrite existing files. true=do overwrite, false=don't overwrite and give a warning. |
Reimplemented in FunctionApproximatorLWR, FunctionApproximatorRBFN, FunctionApproximatorGPR, and FunctionApproximatorRRRFF.
Definition at line 270 of file FunctionApproximator.cpp.
|
protected |
Accessor for FunctionApproximator::model_parameters_.
| [in] | model_parameters | The model parameters of a trained model |
Definition at line 97 of file FunctionApproximator.cpp.
|
friend |
Give boost serialization access to private members.
Definition at line 331 of file FunctionApproximator.hpp.
|
friend |
Print to output stream.
| [in] | output | Output stream to which to write to |
| [in] | function_approximator | Function approximator to write |
Definition at line 244 of file FunctionApproximator.hpp.
 1.8.11
1.8.11