|
DMP_BBO library
|
|
DMP_BBO library
|
GMR (Gaussian Mixture Regression) function approximator. More...
#include <FunctionApproximatorGMR.hpp>


Public Member Functions | |
| FunctionApproximatorGMR (const MetaParametersGMR *const meta_parameters, const ModelParametersGMR *const model_parameters=NULL) | |
| Initialize a function approximator with meta- and model-parameters. More... | |
| FunctionApproximatorGMR (const ModelParametersGMR *const model_parameters) | |
| Initialize a function approximator with model parameters. More... | |
| virtual FunctionApproximator * | clone (void) const |
| Return a pointer to a deep copy of the FunctionApproximator object. More... | |
| void | train (const Eigen::Ref< const Eigen::MatrixXd > &inputs, const Eigen::Ref< const Eigen::MatrixXd > &targets) |
| Train the function approximator with corresponding input and target examples. More... | |
| void | predict (const Eigen::Ref< const Eigen::MatrixXd > &inputs, Eigen::MatrixXd &output) |
| Query the function approximator to make a prediction. More... | |
| void | predictVariance (const Eigen::Ref< const Eigen::MatrixXd > &inputs, Eigen::MatrixXd &variances) |
| Query the function approximator to get the variance of a prediction This function is not implemented by all function approximators. More... | |
| void | predict (const Eigen::Ref< const Eigen::MatrixXd > &inputs, Eigen::MatrixXd &outputs, Eigen::MatrixXd &variances) |
| Query the function approximator to make a prediction, and also to predict its variance. More... | |
| std::string | getName (void) const |
| Get the name of this function approximator. More... | |
| void | predictDot (const Eigen::Ref< const Eigen::MatrixXd > &inputs, Eigen::MatrixXd &outputs, Eigen::MatrixXd &outputs_dot) |
| Query the function approximator to make a prediction and to compute the derivate of that prediction. More... | |
| void | predictDot (const Eigen::Ref< const Eigen::MatrixXd > &inputs, Eigen::MatrixXd &outputs, Eigen::MatrixXd &outputs_dot, Eigen::MatrixXd &variances) |
| Query the function approximator to make a prediction and to compute the derivate of that prediction, and also to predict its variance. More... | |
| void | trainIncremental (const Eigen::Ref< const Eigen::MatrixXd > &inputs, const Eigen::Ref< const Eigen::MatrixXd > &targets) |
| Train the function approximator incrementally with corresponding input and target examples. More... | |
 Public Member Functions inherited from FunctionApproximator Public Member Functions inherited from FunctionApproximator | |
| FunctionApproximator (const MetaParameters *const meta_parameters, const ModelParameters *const model_parameters=NULL) | |
| Initialize a function approximator with meta- and optionally model-parameters. More... | |
| FunctionApproximator (const ModelParameters *const model_parameters) | |
| Initialize a function approximator with model-parameters. More... | |
| void | train (const Eigen::Ref< const Eigen::MatrixXd > &inputs, const Eigen::Ref< const Eigen::MatrixXd > &targets, std::string save_directory, bool overwrite=false) |
| Train the function approximator with corresponding input and target examples (and write results to file). More... | |
| void | reTrain (const Eigen::Ref< const Eigen::MatrixXd > &inputs, const Eigen::Ref< const Eigen::MatrixXd > &targets) |
| Re-train the function approximator with corresponding input and target examples. More... | |
| void | reTrain (const Eigen::Ref< const Eigen::MatrixXd > &inputs, const Eigen::Ref< const Eigen::MatrixXd > &targets, std::string save_directory, bool overwrite=false) |
| Re-train the function approximator with corresponding input and target examples (and write results to file). More... | |
| virtual void | predict (const Eigen::Ref< const Eigen::MatrixXd > &inputs, Eigen::MatrixXd &outputs, std::vector< Eigen::MatrixXd > &variances) |
| Query the function approximator to make a prediction, and also to predict its variance. More... | |
| bool | isTrained (void) const |
| Determine whether the function approximator has already been trained with data or not. More... | |
| int | getExpectedInputDim (void) const |
| The expected dimensionality of the input data. More... | |
| int | getExpectedOutputDim (void) const |
| The expected dimensionality of the output data. More... | |
| void | getSelectableParameters (std::set< std::string > &selected_values_labels) const |
| Return all the names of the parameter types that can be selected. More... | |
| void | setSelectedParameters (const std::set< std::string > &selected_values_labels) |
| Determine which subset of parameters is represented in the vector returned by Parameterizable::getParameterVectorSelected. More... | |
| void | getParameterVectorSelectedMinMax (Eigen::VectorXd &min, Eigen::VectorXd &max) const |
| Get the minimum and maximum of the selected parameters in one vector. More... | |
| int | getParameterVectorSelectedSize (void) const |
| Get the size of the vector of selected parameters, as returned by getParameterVectorSelected(. More... | |
| void | setParameterVectorSelected (const Eigen::VectorXd &values, bool normalized=false) |
| Set all the values of the selected parameters with one vector. More... | |
| void | getParameterVectorSelected (Eigen::VectorXd &values, bool normalized=false) const |
| Get the values of the selected parameters in one vector. More... | |
| void | getParameterVectorMask (const std::set< std::string > selected_values_labels, Eigen::VectorXi &selected_mask) const |
| Get a mask for selecting parameters. More... | |
| int | getParameterVectorAllSize (void) const |
| Get the size of the parameter values vector when it contains all available parameter values. More... | |
| void | getParameterVectorAll (Eigen::VectorXd &values) const |
| Return a vector that returns all available parameter values. More... | |
| void | setParameterVectorAll (const Eigen::VectorXd &values) |
| Set all available parameter values with one vector. More... | |
| UnifiedModel * | getUnifiedModel (void) const |
| Return a representation of this function approximator's model as a unified model. More... | |
| std::string | toString (void) const |
| Returns a string representation of the object. More... | |
| const MetaParameters * | getMetaParameters (void) const |
| Accessor for FunctionApproximator::meta_parameters_. More... | |
| const ModelParameters * | getModelParameters (void) const |
| Accessor for FunctionApproximator::model_parameters_. More... | |
| void | setParameterVectorModifierPrivate (std::string modifier, bool new_value) |
| Turn certain modifiers on or off, see Parameterizable::setParameterVectorModifier(). More... | |
| virtual bool | saveGridData (const Eigen::VectorXd &min, const Eigen::VectorXd &max, const Eigen::VectorXi &n_samples_per_dim, std::string directory, bool overwrite=false) const |
| Generate a grid of inputs, and output the response of the basis functions and line segments for these inputs. More... | |
 Public Member Functions inherited from Parameterizable Public Member Functions inherited from Parameterizable | |
| virtual | ~Parameterizable (void) |
| Destructor. | |
| virtual void | getParameterVectorSelectedNormalized (Eigen::VectorXd &values) const |
| Get the normalized values of the selected parameters in one vector. More... | |
| void | getParameterVectorSelectedMinMax (Eigen::VectorXd &min, Eigen::VectorXd &max) const |
| Get the minimum and maximum of the selected parameters in one vector. More... | |
| void | getParameterVectorAllMinMax (Eigen::VectorXd &min, Eigen::VectorXd &max) const |
| Get the minimum and maximum values of the current parameter vector. More... | |
| void | getParameterVectorSelectedRanges (Eigen::VectorXd &ranges) const |
| Get the ranges of the selected parameters, i.e. More... | |
| virtual void | setParameterVectorSelectedNormalized (const Eigen::VectorXd &values) |
| Set all the values of the selected parameters with one vector of normalized values. More... | |
| void | setSelectedParametersOne (std::string selected) |
| Set the parameters that are currently selected. More... | |
| void | setParameterVectorModifier (std::string modifier, bool new_value) |
| Turn certain modifiers on or off. More... | |
| void | setVectorLengthsPerDimension (const Eigen::VectorXi &lengths_per_dimension) |
| The vector (VectorXd) with parameter values can be split into different parts (as vector<VectorXd>; this function specifices the length of each sub-vector. More... | |
| Eigen::VectorXi | getVectorLengthsPerDimension (void) const |
| Get the specified length of each vector in each dimension. More... | |
| void | getParameterVectorSelected (std::vector< Eigen::VectorXd > &values, bool normalized=false) const |
| Get the values of the selected parameters in one vector. More... | |
| void | setParameterVectorSelected (const std::vector< Eigen::VectorXd > &values, bool normalized=false) |
| Set all the values of the selected parameters with a vector of vectors. More... | |
Protected Member Functions | |
| void | kMeansInit (const Eigen::MatrixXd &data, std::vector< Eigen::VectorXd > &means, std::vector< double > &priors, std::vector< Eigen::MatrixXd > &covars, int n_max_iter=1000) |
| Initialize Gaussian for EM algorithm using k-means. More... | |
| void | firstDimSlicingInit (const Eigen::MatrixXd &data, std::vector< Eigen::VectorXd > &means, std::vector< double > &priors, std::vector< Eigen::MatrixXd > &covars) |
| Initialize Gaussian for EM algorithm using a same-size slicing on the first dimension (method used in Calinon GMR implementation). More... | |
| void | expectationMaximization (const Eigen::MatrixXd &data, std::vector< Eigen::VectorXd > &means, std::vector< double > &priors, std::vector< Eigen::MatrixXd > &covars, int n_max_iter=50) |
| EM algorithm. More... | |
| void | expectationMaximizationIncremental (const Eigen::MatrixXd &data, std::vector< Eigen::VectorXd > &means, std::vector< double > &priors, std::vector< Eigen::MatrixXd > &covars, int &n_observations, int n_max_iter=50) |
| EM algorithm Incremental. More... | |
 Protected Member Functions inherited from FunctionApproximator Protected Member Functions inherited from FunctionApproximator | |
| void | setModelParameters (ModelParameters *model_parameters) |
| Accessor for FunctionApproximator::model_parameters_. More... | |
| FunctionApproximator (void) | |
| Default constructor. More... | |
Static Protected Member Functions | |
| static double | normalPDF (const Eigen::VectorXd &mu, const Eigen::MatrixXd &covar, const Eigen::VectorXd &input) |
| The probability density function (PDF) of the multi-variate normal distribution. More... | |
| static double | normalPDFDamped (const Eigen::VectorXd &mu, const Eigen::MatrixXd &covar, const Eigen::VectorXd &input) |
| The damped probability density function (PDF) of the multi-variate normal distribution. More... | |
Friends | |
| class | boost::serialization::access |
| Give boost serialization access to private members. More... | |
Additional Inherited Members | |
 Static Public Member Functions inherited from FunctionApproximator Static Public Member Functions inherited from FunctionApproximator | |
| static void | generateInputsGrid (const Eigen::VectorXd &min, const Eigen::VectorXd &max, const Eigen::VectorXi &n_samples_per_dim, Eigen::MatrixXd &inputs_grid) |
| Generate a input samples that lie on a grid (much like Matlab's meshgrid) For instance, if min = [2 6], and max = [3 8], and n_samples_per_dim = [3 5] then this function first makes linearly spaces samples along each dimension, e.g. More... | |
GMR (Gaussian Mixture Regression) function approximator.
Definition at line 44 of file FunctionApproximatorGMR.hpp.
| FunctionApproximatorGMR | ( | const MetaParametersGMR *const | meta_parameters, |
| const ModelParametersGMR *const | model_parameters = NULL |
||
| ) |
Initialize a function approximator with meta- and model-parameters.
| [in] | meta_parameters | The training algorithm meta-parameters |
| [in] | model_parameters | The parameters of the trained model. If this parameter is not passed, the function approximator is initialized as untrained. In this case, you must call FunctionApproximator::train() before being able to call FunctionApproximator::predict(). Either meta_parameters XOR model-parameters can passed as NULL, but not both. |
Definition at line 51 of file FunctionApproximatorGMR.cpp.

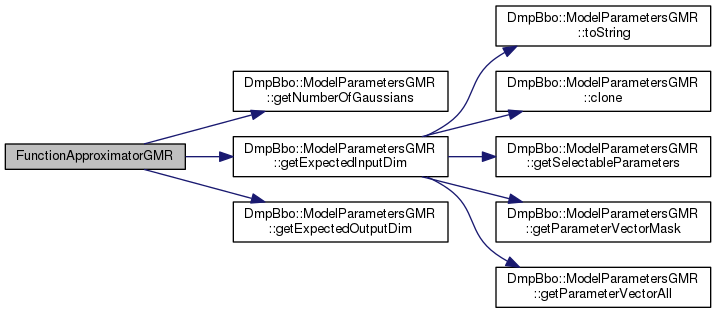
| FunctionApproximatorGMR | ( | const ModelParametersGMR *const | model_parameters | ) |
Initialize a function approximator with model parameters.
| [in] | model_parameters | The parameters of the (previously) trained model. |
Definition at line 65 of file FunctionApproximatorGMR.cpp.
|
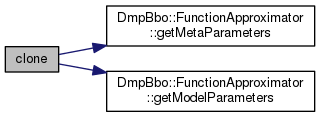
virtual |
Return a pointer to a deep copy of the FunctionApproximator object.
Implements FunctionApproximator.
Definition at line 93 of file FunctionApproximatorGMR.cpp.
|
virtual |
Train the function approximator with corresponding input and target examples.
| [in] | inputs | Input values of the training examples |
| [in] | targets | Target values of the training examples |
Implements FunctionApproximator.
Definition at line 102 of file FunctionApproximatorGMR.cpp.

|
virtual |
Query the function approximator to make a prediction.
| [in] | inputs | Input values of the query |
| [out] | outputs | Predicted output values |
Implements FunctionApproximator.
Definition at line 323 of file FunctionApproximatorGMR.cpp.
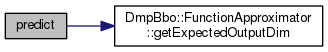
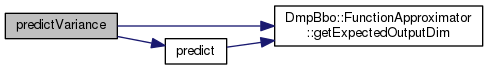
|
virtual |
Query the function approximator to get the variance of a prediction This function is not implemented by all function approximators.
Therefore, the default implementation fills outputs with 0s.
| [in] | inputs | Input values of the query (n_samples X n_dims_in) |
| [out] | variances | Predicted variances for the output values (n_samples X n_dims_out). Note that if the output has a dimensionality>1, these variances should actuall be covariance matrices (use function predict(const Eigen::Ref<const Eigen::MatrixXd>& inputs, Eigen::MatrixXd& outputs, std::vector<Eigen::MatrixXd>& variances) to get the full covariance matrices). So for an output dimensionality of 1 this function works fine. For dimensionality>1 we return only the diagional of the covariance matrix, which may not always be what you want. |
Reimplemented from FunctionApproximator.
Definition at line 331 of file FunctionApproximatorGMR.cpp.
|
virtual |
Query the function approximator to make a prediction, and also to predict its variance.
| [in] | inputs | Input values of the query (n_samples X n_dims_in) |
| [out] | outputs | Predicted output values (n_samples X n_dims_out) |
| [out] | variances | Predicted variances for the output values (n_samples X n_dims_out). Note that if the output has a dimensionality>1, these variances should actuall be covariance matrices (use function predict(const Eigen::Ref<const Eigen::MatrixXd>& inputs, Eigen::MatrixXd& outputs, std::vector<Eigen::MatrixXd>& variances) to get the full covariance matrices). So for an output dimensionality of 1 this function works fine. For dimensionality>1 we return only the diagional of the covariance matrix, which may not always be what you want. |
Reimplemented from FunctionApproximator.
Definition at line 339 of file FunctionApproximatorGMR.cpp.

|
inlinevirtual |
Get the name of this function approximator.
Implements FunctionApproximator.
Definition at line 72 of file FunctionApproximatorGMR.hpp.

|
protected |
Initialize Gaussian for EM algorithm using k-means.
| [in] | data | A data matrix (n_exemples x (n_in_dim + n_out_dim)) |
| [out] | means | A list (std::vector) of n_gaussian non initiallized means (n_in_dim + n_out_dim) |
| [out] | priors | A list (std::vector) of n_gaussian non initiallized priors |
| [out] | covars | A list (std::vector) of n_gaussian non initiallized covariance matrices ((n_in_dim + n_out_dim) x (n_in_dim + n_out_dim)) |
| [in] | n_max_iter | The maximum number of iterations |
Definition at line 696 of file FunctionApproximatorGMR.cpp.
|
protected |
Initialize Gaussian for EM algorithm using a same-size slicing on the first dimension (method used in Calinon GMR implementation).
Particulary suited when input is 1-D and data distribution is uniform over input dimension
| [in] | data | A data matrix (n_exemples x (n_in_dim + n_out_dim)) |
| [out] | means | A list (std::vector) of n_gaussian non initiallized means (n_in_dim + n_out_dim) |
| [out] | priors | A list (std::vector) of n_gaussian non initiallized priors |
| [out] | covars | A list (std::vector) of n_gaussian non initiallized covariance matrices ((n_in_dim + n_out_dim) x (n_in_dim + n_out_dim)) |
Definition at line 647 of file FunctionApproximatorGMR.cpp.
|
protected |
EM algorithm.
| [in] | data | A (n_exemples x (n_in_dim + n_out_dim)) data matrix |
| [in,out] | means | A list (std::vector) of n_gaussian means (vector of size (n_in_dim + n_out_dim)) |
| [in,out] | priors | A list (std::vector) of n_gaussian priors |
| [in,out] | covars | A list (std::vector) of n_gaussian covariance matrices ((n_in_dim + n_out_dim) x (n_in_dim + n_out_dim)) |
| [in] | n_max_iter | The maximum number of iterations |
Definition at line 778 of file FunctionApproximatorGMR.cpp.

|
protected |
EM algorithm Incremental.
| [in] | data | A (n_exemples x (n_in_dim + n_out_dim)) data matrix |
| [in,out] | means | A list (std::vector) of n_gaussian means (vector of size (n_in_dim + n_out_dim)) |
| [in,out] | priors | A list (std::vector) of n_gaussian priors |
| [in,out] | covars | A list (std::vector) of n_gaussian covariance matrices ((n_in_dim + n_out_dim) x (n_in_dim + n_out_dim)) |
| [in,out] | n_observations | Number of observations |
| [in] | n_max_iter | The maximum number of iterations |
Definition at line 883 of file FunctionApproximatorGMR.cpp.

|
staticprotected |
The probability density function (PDF) of the multi-variate normal distribution.
| [in] | mu | The mean of the normal distribution |
| [in] | covar | The covariance matrix of the normal distribution |
| [in] | input | The input data vector for which the PDF will be computed. |
Definition at line 283 of file FunctionApproximatorGMR.cpp.
|
staticprotected |
The damped probability density function (PDF) of the multi-variate normal distribution.
| [in] | mu | The mean of the normal distribution |
| [in] | covar | The covariance matrix of the normal distribution |
| [in] | input | The input data vector for which the PDF will be computed. |
Definition at line 297 of file FunctionApproximatorGMR.cpp.
| void predictDot | ( | const Eigen::Ref< const Eigen::MatrixXd > & | inputs, |
| Eigen::MatrixXd & | outputs, | ||
| Eigen::MatrixXd & | outputs_dot | ||
| ) |
Query the function approximator to make a prediction and to compute the derivate of that prediction.
| [in] | inputs | Input values of the query |
| [out] | outputs | Predicted output values |
| [out] | outputs_dot | Predicted derivate values |
Definition at line 479 of file FunctionApproximatorGMR.cpp.
| void predictDot | ( | const Eigen::Ref< const Eigen::MatrixXd > & | inputs, |
| Eigen::MatrixXd & | outputs, | ||
| Eigen::MatrixXd & | outputs_dot, | ||
| Eigen::MatrixXd & | variances | ||
| ) |
Query the function approximator to make a prediction and to compute the derivate of that prediction, and also to predict its variance.
| [in] | inputs | Input values of the query (n_samples X n_dims_in) |
| [out] | outputs | Predicted output values (n_samples X n_dims_out) |
| [out] | outputs_dot | Predicted derivate values |
| [out] | variances | Predicted variances for the output values (n_samples X n_dims_out). Note that if the output has a dimensionality>1, these variances should actuall be covariance matrices (use function predict(const Eigen::Ref<const Eigen::MatrixXd>& inputs, Eigen::MatrixXd& outputs, std::vector<Eigen::MatrixXd>& variances) to get the full covariance matrices). So for an output dimensionality of 1 this function works fine. For dimensionality>1 we return only the diagional of the covariance matrix, which may not always be what you want. |
Definition at line 487 of file FunctionApproximatorGMR.cpp.

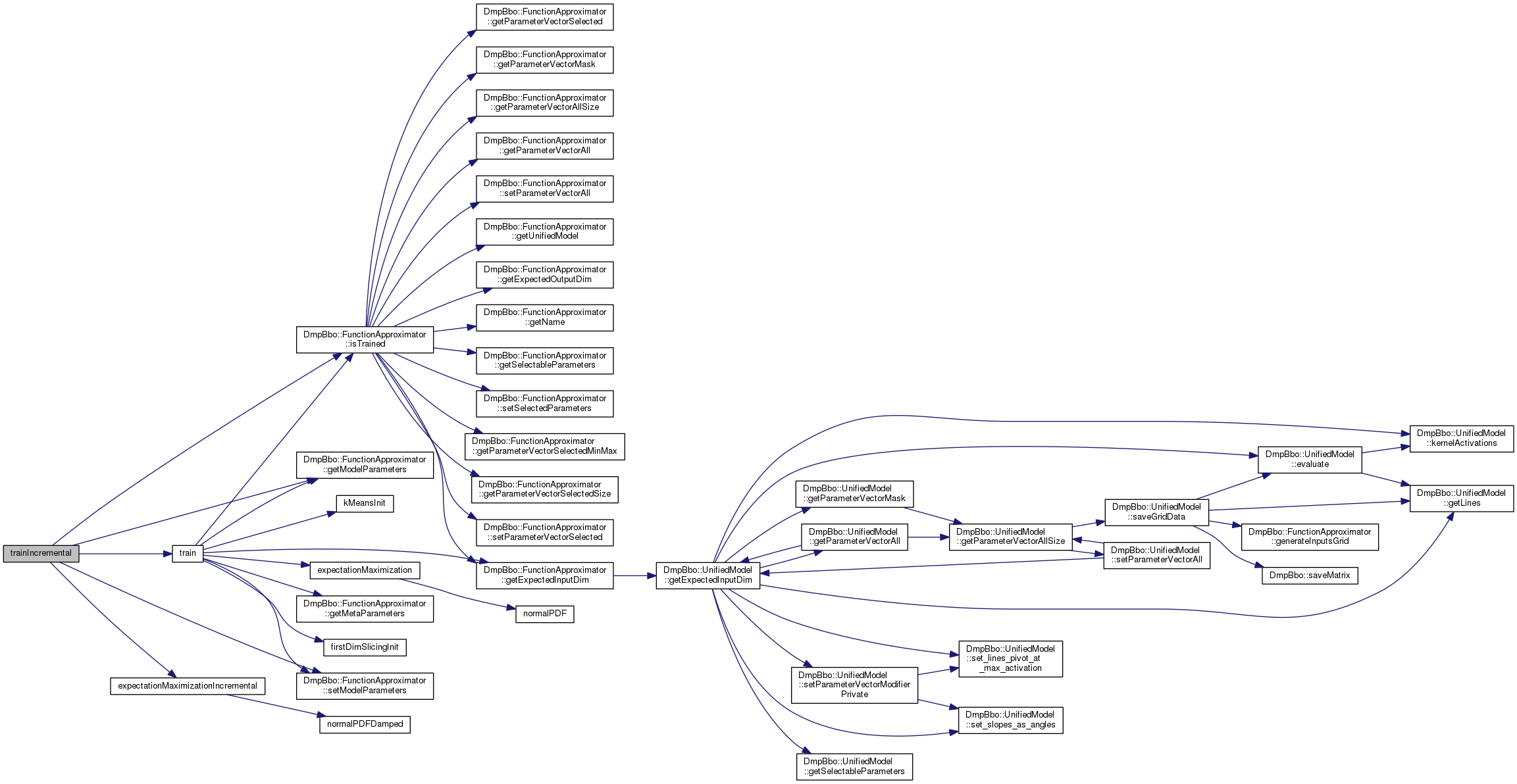
| void trainIncremental | ( | const Eigen::Ref< const Eigen::MatrixXd > & | inputs, |
| const Eigen::Ref< const Eigen::MatrixXd > & | targets | ||
| ) |
Train the function approximator incrementally with corresponding input and target examples.
| [in] | inputs | Input values of the training examples |
| [in] | targets | Target values of the training examples |
Definition at line 211 of file FunctionApproximatorGMR.cpp.
|
friend |
Give boost serialization access to private members.
Definition at line 177 of file FunctionApproximatorGMR.hpp.
 1.8.11
1.8.11